Who we are
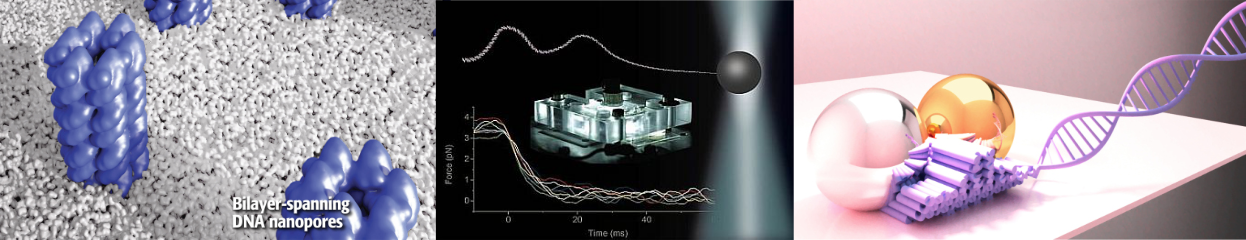
We are a group of scientists at the Cavendish Lab, University of Cambridge, UK. Our research is focused on understanding transport processes through membranes for biosensing applications.
Since the pandemic we are mainly interested in understanding RNA, its structure and its relation to biology and disease. More details on our current and past research interests can be found here. Since the start, the lab aims to achieve a maximum level of control over all parameters in our experiments. Our main technique remains resistive-pulse sensing with nanopores especially in combination with DNA and now RNA nanotechnology
Our interdisciplinary team combines researchers with expertise in physics, engineering, physical chemistry, biochemistry/biology, and micro- and nanofabrication.
In case you are interested in working with us, please get in touch with Ulrich by email: ufk20 (at) cam.ac.uk.
We gratefully acknowledge funding of our work from various sources including:
 |
 |
 |
||
 |
 |
 |
News
4/05/2026 Single-molecule nanoarray for short RNA
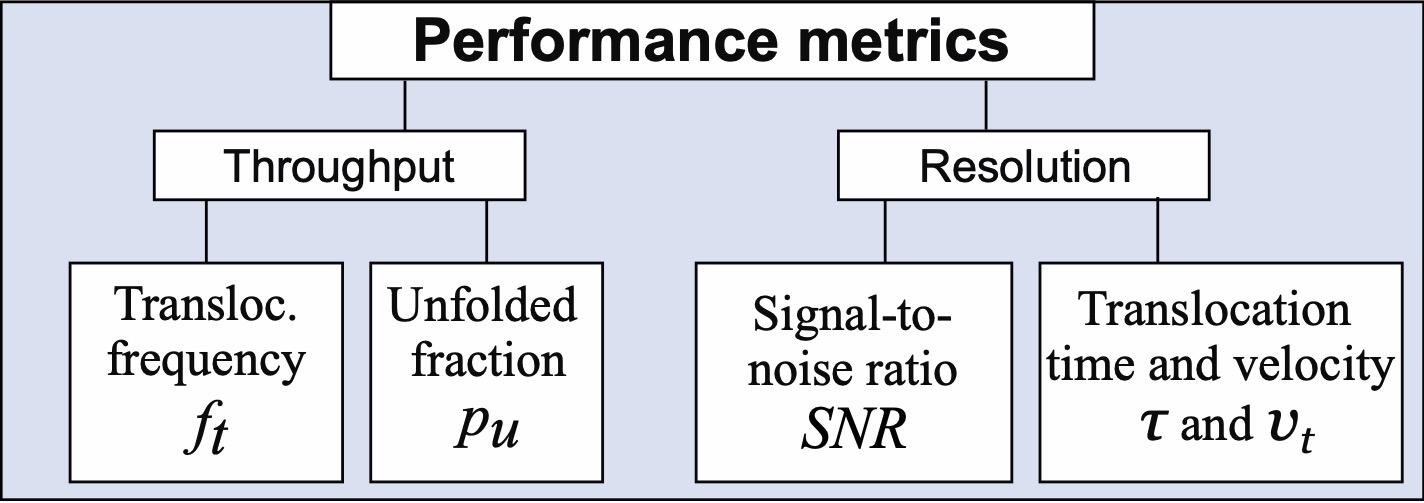
24/03/2026 Our guide for optimal nanopore sensing for nucleic acids.
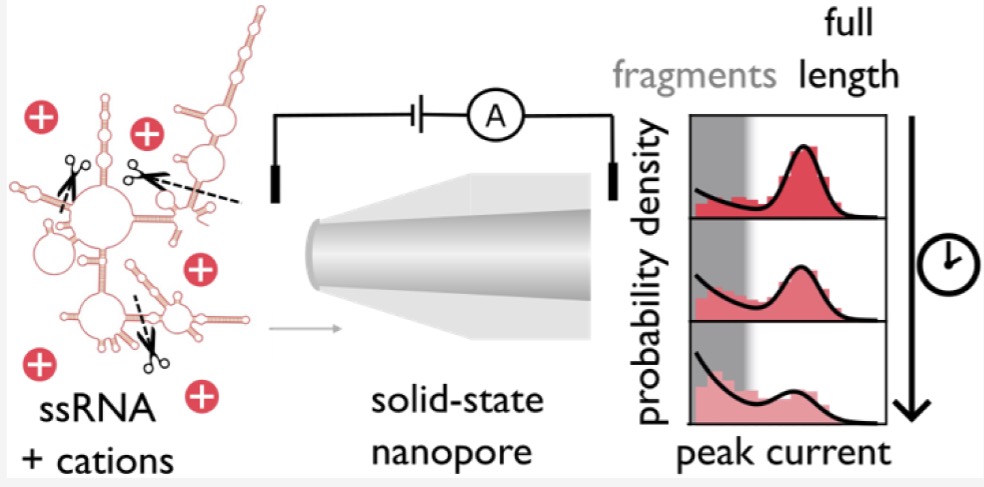
7/11/2025 Glass nanopore biosensing for quantification of RNA degradation.
7/11/2025 Raluca wins Best Poster Award at Sensors Day 2025.

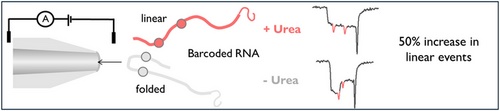
16/10/2025 Urea changes translocation of RNA:DNA hybrids.